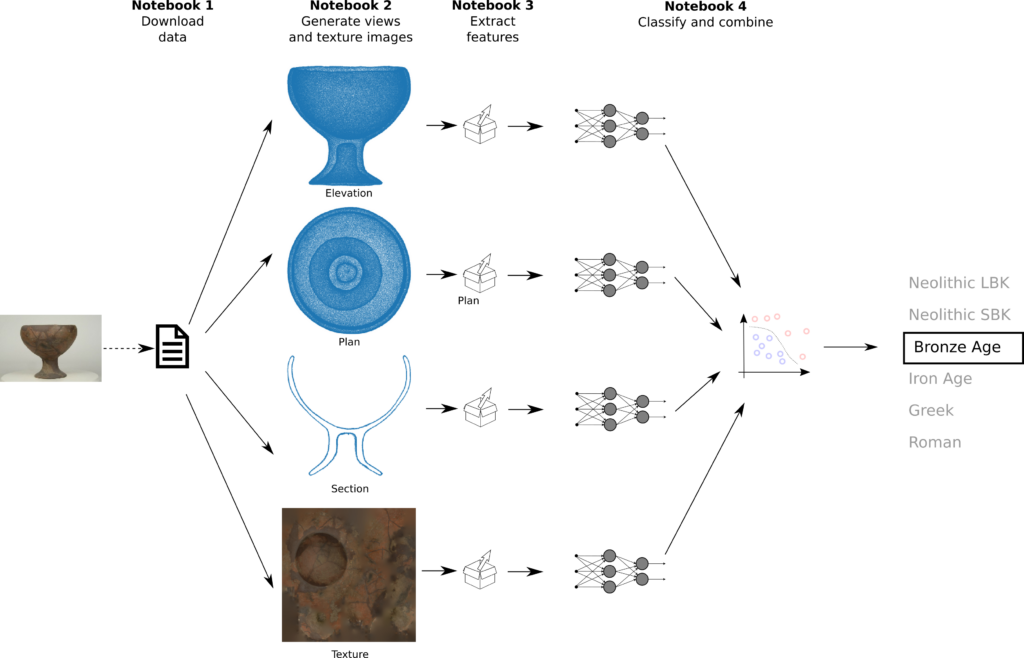
There are important potential applications for a machine learning system than can classify plants. For example, current research on crop protection is using machine learning for precision weed and plant disease detection. The importance of moving away from traditional pesticide-based crop protection methods cannot be overstated, as demonstrated by the alarming rate at which flowering plants are evolving away from insect pollination.
I obtained a dataset of plant leaf images (Hussain, 2023) and compared three machine learning algorithms for classification: multilayer perceptron, random forest and support vector machine. The classifiers’ accuracy vary between 74% and 84% .
The Jupyter notebook with the full Python code can be accessed on Kaggle.
![Rendered by QuickLaTeX.com \[D = 1 - \sum_{i=1}^{S}p_i^2\]](http://biodiv.smultron.org/wp-content/ql-cache/quicklatex.com-55f32c820d77a3a7dc38c88992946e12_l3.png)